Services on Demand
Article
Indicators
Related links
-
 Cited by Google
Cited by Google -
 Similars in Google
Similars in Google
Share
SAMJ: South African Medical Journal
On-line version ISSN 2078-5135
Print version ISSN 0256-9574
SAMJ, S. Afr. med. j. vol.104 n.8 Pretoria Aug. 2014
RESEARCH
National sentinel site surveillance for antimicrobial resistance in Klebsiella pneumoniae isolates in South Africa, 2010 - 2012
O PerovicI; A Singh-MoodleyII; A DuséIII; C BamfordIV; G ElliottV; K Swe Swe-HanVI; R KularatneVII; W LowmanVIII; A WhitelawIX; T NanaX; J WadulaXI; R LekalakalaXII; A SaifXIII; M Fortuin De-SmidtIIV; E MaraisXV
IMD, DTM&H, FCPath (SA) (Microbiol), MMed (Microbiol). Centre for Opportunistic, Tropical and Hospital Infections, National Institute for Communicable Diseases, National Health Laboratory Service, Johannesburg, South Africa. Department of Clinical Microbiology and Infectious Diseases, School of Pathology of the University of the Witwatersrand and National Health Laboratory Service, Johannesburg, South Africa
IIBSc, BMedSc (Hons), MMedSc, PhD; Centre for Opportunistic, Tropical and Hospital Infections, National Institute for Communicable Diseases, National Health Laboratory Service, Johannesburg, South Africa. Department of Clinical Microbiology and Infectious Diseases, School of Pathology of the University of the Witwatersrand and National Health Laboratory Service, Johannesburg, South Africa
IIIMB BCh, DTM&H, MSc Med, MMed, FCPath (SA) (Microbiol); Department of Clinical Microbiology and Infectious Diseases, School of Pathology of the University of the Witwatersrand and National Health Laboratory Service, Johannesburg, South Africa
IVMB ChB, DCH, MPhil, FCPath (SA) (Microbiol), MMed; Division of Medical Microbiology, Department of Clinical Laboratory Sciences, Faculty of Health Sciences, University of Cape Town and National Health Laboratory Service, Groote Schuur Hospital, Cape Town, South Africa
VMB ChB, FCPath (SA) (Microbiol), MMed (Microbiol), MBA; InterSystems, Sandton, Johannesburg, South Africa
VIFCPath (SA) (Microbiol), MMed (Microbiol), DTM&H, PDIC, MBBS; Department of Medical Microbiology, Inkosi Albert Luthuli Hospital, Durban, South Africa, and Microbiology Department, National Health Laboratory Service, University of KwaZulu-Natal, Durban
VIIBSc, MB ChB, MSc Med (Microbiol), FCPath (SA) (Microbiol), DTM&H, Dip HIV Man (SA); Department of Clinical Microbiology and Infectious Diseases, School of Pathology of the University of the Witwatersrand and National Health Laboratory Service, Johannesburg, South Africa
VIIIMB BCh, MMed, FCPath (SA) (Microbiol); Department of Clinical Microbiology and Infectious Diseases, School of Pathology of the University of the Witwatersrand and National Health Laboratory Service, Johannesburg, South Africa. Wits University Donald Gordon Medical Centre, Parktown, Johannesburg, South Africa
IXMB BCh, MSc, FCPath (SA) (Microbiol); Division of Medical Microbiology, Faculty of Medicine and Health Sciences, Stellenbosch University and National Health Laboratory Service, Tygerberg Hospital, Cape Town, South Africa
XMB BCh, DTM&H, FCPath (SA) (Microbiol), MMed; Department of Clinical Microbiology and Infectious Diseases, School of Pathology of the University of the Witwatersrand and National Health Laboratory Service, Johannesburg, South Africa
XIMD, FCPath (SA) (Microbiol), DTM&H, PDIC; Department of Clinical Microbiology and Infectious Diseases, School of Pathology of the University of the Witwatersrand and National Health Laboratory Service, Johannesburg, South Africa
XIIBSc, MB ChB, MMed, DTM&H, PDIC; Department of Medical Microbiology, University of Pretoria and Tshwane Academic Division, National Health Laboratory Service, Pretoria, South Africa
XIIIMSc Med (Microbiol); Centre for Opportunistic, Tropical and Hospital Infections, National Institute for Communicable Diseases, National Health Laboratory Service, Johannesburg, South Africa
XIVBSc, BSc (Hons), MB ChB, MSc; Centre for Opportunistic, Tropical and Hospital Infections, National Institute for Communicable Diseases, National Health Laboratory Service, Johannesburg, South Africa
XVPhD. Centre for Opportunistic, Tropical and Hospital Infections, National Institute for Communicable Diseases, National Health Laboratory Service, Johannesburg, South Africa
ABSTRACT
BACKROUND: The increasing rates of antimicrobial resistance observed in the nosocomial pathogen Klebsiella pneumoniae are of major public health concern worldwide.
OBJECTIVES: To describe the antibiotic susceptibility profiles of K. pneumoniae isolates from bacteraemic patients submitted by sentinel laboratories in five regions of South Africa from mid-2010 to mid-2012. Molecular methods were used to detect the most commonly found extended-spectrum beta-lactamase (ESBL) and carbapenemase resistance genes.
METHODS: Thirteen academic centres serving the public healthcare sector in Gauteng, KwaZulu-Natal, Free State, Limpopo and Western Cape provinces submitted K. pneumoniae isolates from patients with bloodstream infections. Vitek 2 and MicroScan instruments were used for organism identification and susceptibility testing. Multiplex polymerase chain reactions (PCRs) were used to detect blaCTX-M, blaSHV and blaTEM genes in a proportion of the ESBL isolates. All isolates exhibiting reduced susceptibility to carbapenems were PCR tested for blaKPC and blaNDM-1 resistance genes.
RESULTS: Overall, 68.3% of the 2 774 isolates were ESBL-positive, showing resistance to cefotaxime, ceftazidime and cefepime. Furthermore, 46.5% of all isolates were resistant to ciprofloxacin and 33.1% to piperacillin-tazobactam. The major ESBL genes were abundantly present in the sample analysed. Most isolates (95.5%) were susceptible to the carbapenems tested, and no isolates were positive for blaKPC or blaNDM1 There was a trend towards a decrease in susceptibility to most antibiotics.
CONCLUSION: The high proportion of ESBL-producing K. pneumoniae isolates observed, and the prevalence of ESBL genes, are of great concern. Our findings represent a baseline for further surveillance in SA, and can be used for policy and treatment decisions.
The increasing prevalence of serious hospital-and community-acquired infections is of great concern, with patients dying as a result of emerging antimicrobial resistance. Antibiotic-resistant organisms are widespread globally, both in developed and developing countries.[1] These organisms are a major threat to public health, reduce the effectiveness of empiric antimicrobial treatment options, and increase morbidity, mortality and healthcare expenditure. A national surveillance system for antimicrobial resistance is essential to establish a baseline of the extent of the problem, to follow trends of resistance, and to form the basis for recommendations of appropriate antimicrobial use to clinicians and other healthcare providers. A national system also controls for differences in case selection, data management and demographic descriptions of regional populations. To meet this need, a national laboratory-based antimicrobial resistance surveillance system for nosocomial pathogens was established in 2010, which included Klebsiella pneumoniae as a sentinel organism by which to monitor resistance.
K. pneumoniae is an important noso-comial pathogen, with the highest prevalence of resistance to third- and fourth-generation cephalosporins among the Enterobacteriaceae. The spread of class A or group 2be extended-spectrum beta-lactamases (ESBLs) in Enterobacteriaceae is of public health concern. The most frequently detected and clinically important ESBLs belong to the TEM, SHV, and CTX-M families, and K. pneumoniae commonly produces all three groups of enzymes.[2] In the past decade, CTX-M enzymes have emerged as the most prevalent type. There are more than 100 different types, which can be broadly divided into five groups based on their amino-acid identities: CTX-M1, CTX-M2, CTX-M8, CTX-M9 and CTX-M25. Enzymes are characterised by epidemiological differences, and some have even been shown to spread beyond the hospital environment into the community.[2] Of further public health concern are the recent emergence of K. pneumoniae strains capable of producing carbapenem-hydrolysing enzymes and the apparent ease of spread of resistance mechanisms by mobile genetic elements.[2]
There are few published reports on national antimicrobial resistance rates of K. pneumoniae in South Africa (SA), with data primarily emanating from the private sector,[3] regional studies with limited numbers of local isolates[1] or certain clinical settings.[4] To date, molecular data on ESBL K. pneumoniae in SA are based on detailed studies of small populations. Essack et al.[5] examined blaSHV and blaTEM genes in 25 isolates, and found a high degree of diversity in terms of plasmids present, genes detected and the combination of genes within isolates. In a study of ESBL K. pneumoniae from seven countries in 2003, blaTEM-10, blaTEM-12, bla TEM-63VariousblaSHV-2 and blaSHV-5 types, blaCTX-M2 and blaCTX-M3 genes were detected in 27 SA isolates.[61] A more recent study on 53 ESBL clinical pathogens from Pretoria detected blaTEM, blaSHV and blaCTX-M-1 genes, although the contribution of the 31 K. pneumoniae isolates in this group was not specified.[7] This diversity in enzyme production and the prolific nature of K. pneumoniae as a nosocomial pathogen highlight the need for further investigation into the genes responsible for these enzymes in the SA setting.
In this study, we determined the antibiotic susceptibility profiles of K. pneumo-niae isolates from bacteraemic patients with positive blood cultures from sentinel sites representing five SA regions. We used national laboratory-based surveillance data to characterise the third- and fourth-generation cephalosporin-resistant and carbapenem-resistant phenotypes and genotypes of K. pneumoniae from 2010 to 2012.
Methods
Patient selection
Thirteen academic centres serving the public healthcare sector in SA were included, and participation was voluntary. The sites represented regions in Gauteng, KwaZulu-Natal, Free State, Limpopo and Western Cape provinces. Isolates of K. pneumoniae from patients with bloodstream infections were submitted, with a 3-week exclusion thereafter to avoid duplicate isolates of the same organism.
Phenotypic methods
K. pneumoniae isolates were submitted on Dorset transport media. Organism identification was confirmed using the Vitek 2 GN card (Biomerieux, France). Susceptibility testing and determination of ESBL phenotype was performed on the MicroScan Walkaway (Siemens Healthcare Diagnostics, USA), using NM37 panels. Categorical results and susceptibility profile of each antimicrobial agent tested were based on 2012 Clinical Laboratory Standards Institute (CLSI) interpretative criteria,[81 the European Committee on Antimicrobial Susceptibility Testing guidelines[9] and/ or the MicroScan recommendations. The MIC50 and MIC90 (minimum inhibitory concentrations needed to inhibit the growth of 50% and 90% of organisms, respectively) were determined. The Agresti-Coull interval method was used to calculate confidence intervals, and a χ2 test was performed to analyse trends in antibiotic susceptibility. A p-value of <0.05 was deemed statistically significant.
Genotypic methods
Two hundred and seventy ESBL-producing isolates were randomly selected (approximately 14% per region) and screened for the presence of ESBL genes. DNA was extracted from half a loop (~2 mm in diameter) of bacterial culture. This was resuspended in 400 µl tris-ethylenediaminetetraacetic acid buffer (pH 8.0), vortexed briefly, heated at 95°C for 25 minutes and pelleted by centrifugation. The supernatant was aliquoted and stored at -70°C for further use.
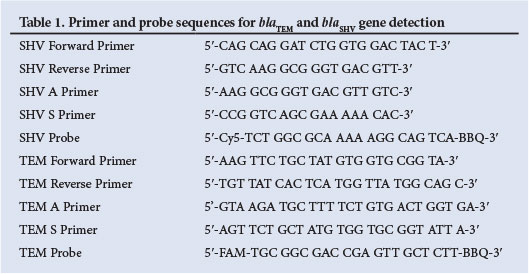
The LightCycler 480 instrument (Roche Applied Science, Germany) and LightCycler 480 Probes Master kit (Roche Diagnostics, USA) were used for real-time polymerase chain reaction (PCR). The blaTEM and blaSHV genes were amplified by multiplex real-time PCR using the primers shown in Table 1. The primers were selected for specificity by GenBank comparisons and PCR products from control strains were sequenced. The limit of detection was determined to be 750 colony-forming units (cfu)/ml for blaTEM and 4 000 cfu/ml for blaSHV. The reaction conditions were 0.5 µM primers, 0.2 µM probes, denaturation for 95°C for 5 minutes, and then 45 cycles of 95°C for 10 seconds, 55°C for 30 seconds and 72°C for 1 second. The blaCTX-M PCR was a multiplex assay targeting blaCTX-M groups M1 and M2-9 using primer and probe sequences as described previously.[10] Detection of blaNDM1 and blaKpc genes was performed on isolates with reduced susceptibility to carbapenems as described previously.[11] Reduced susceptibility to carbapenems was defined according to the CLSI guidelines of 2012.[8] Strains from the American Type Culture Collection (ATCC) or the National Culture Collection Laboratory of the National Institute for Communicable Diseases were used as positive controls.

Results
A total of 2 774 discrete K. pneumoniae isolates were received and included in the antimicrobial susceptibility testing over the 3-year period. The majority of isolates demonstrated an ESBL phenotype (68.3%) with marked resistance to third- and fourth-generation cephalosporins. The distribution of isolates according to province, and ESBL rates, are shown in Table 2. A breakdown of susceptibility by agent for the 3-year period is presented in Fig. 1.


Reduced carbapenem susceptibility was noted in only 124 (4.5%) of isolates (Table 2). The carbapenemase genes blaNDM-1 and blaKPC were not detected in any of the 124 isolates and the presence of other carbapenemase-producing genes was not investigated.
Susceptibility to aminoglycosides over the 3-year period was variable, with 95.3% susceptible to amikacin but only 31.1% and 31.3% susceptible to tobramycin and gentamicin, respectively (Fig. 1). Susceptibility to levofloxacin (75.2%) was higher than to ciprofloxacin (53.5%). Sixty-seven (66.9%) per cent of all isolates were susceptible to piperacillin-tazobactam (MIC50 = 8 µg/ml) for the 3 years, and susceptibility to cefoxitin was high at 87.4% (Table 3).
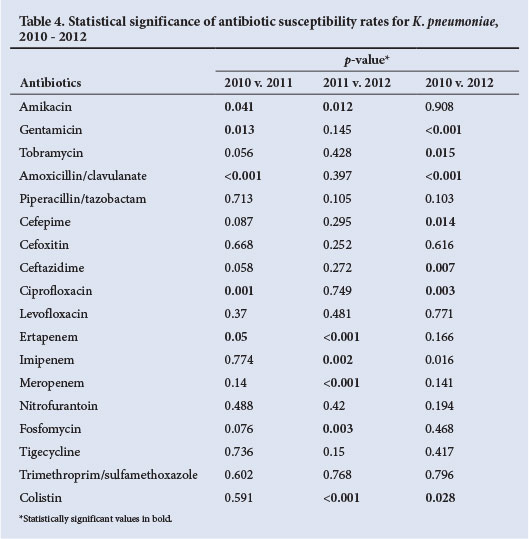
Trend analysis was performed on the isolates, comparing the susceptibility rates between 2010, 2011 and 2012 (Table 4). Small but statistically significant declines in susceptibility rates were noted for many antibiotics, including amoxicillin/ clavulanate, tobramycin, gentamicin, cipro-floxacin, ceftazidime and cefotaxime.
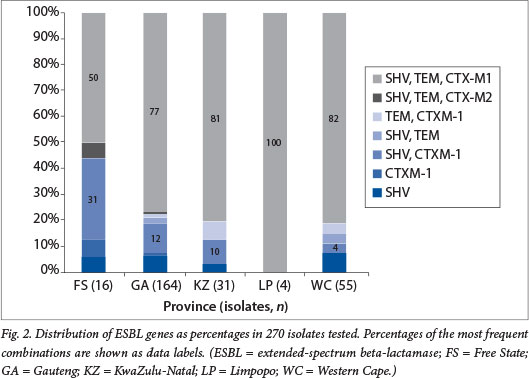
Molecular characterisation was performed on 270 ESBL-positive isolates. All phenotypically ESBL-positive isolates were confirmed to possess one or more of the blaCTX-M, blaSHV and blaTEM genes, with 93.0% of the isolates tested expressing more than one resistance gene (Fig. 2).
Discussion
Gauteng contributed the majority of isolates in the study (62.6%), probably because it is the most populous province with the largest academic centres. The high percentage of bacteraemic K. pneumoniae that were resistant to third-generation cephalosporins (68.3%) is of serious public health concern. Rates of resistance in a 2006 study from SA private laboratories showed 52% resistance to cefuroxime, 46% to ceftriaxone and 44% to cefepime.[31 A study of resistance at seven public sector hospitals between 2010 and 2012 reported an overall level of ESBL detection of 65%.[12] Our study shows a similar rate (68.3%) of resistance to extended-spectrum cephalosporins. The difference between the private and public sectors may indicate a dramatic increase in development of resistance in the interim, and/or a difference in sampling and resistance patterns between these sectors. Also, because they are referral centres, the academic centres that submitted isolates in this study potentially contributed a disproportionate number of patients who were more likely to harbour ESBL isolates. Globally, reports show a trend towards an increase in resistance of K. pneumoniae to third-generation cephalosporins.[13] While rates of resistance are low in some countries, e.g. 11.5% in 2010 in the USA,[13] higher rates have been observed in other regions.[14]
This laboratory-based surveillance also underl ines the endemic distribution of all three major groups of ESBL genes throughout five regions of SA. The almost identical susceptibility pattern to cefotaxime, cef-tazidime and cefepime suggests that there are multiple copies of several ESBL genes in the sample, which is supported by the genotypic results. A limitation of the study is that the genotypic data represent only 14% of K. pneumoniae isolates from selected academic or referral laboratories, and may not be fully representative at a national level. The phenotypic data support the genotypic data, however, being fairly representative of all isolates sampled in this survey. These results are also a first indication of the extensive prevalence of blaSHV, blaTEM and blaCTXM genes in geographically distinct regions of SA.
Susceptibility to carbapenems was high, with only 4.5% of isolates showing reduced susceptibility. However, carbapenem-resis-tant Enterobacteriaceae have been described in SA,[15] and levels of carbapenem resistance need to be monitored closely. It must also be noted that the breakpoints for ertapenem have since been revised by the CLSI.[81 A limitation of this study is that we only tested for blaNDM-1 and blaKPC genes and did not investigate for other potential mechanisms of carbapenem resistance.
For piperacillin-tazobactam, the MIC50 was within the susceptible breakpoint while the MIC90 was above the resistant breakpoint. Overall susceptibility was 66.9%, and piperacillin-tazobactam may therefore be a potentially useful agent for treatment. However, there is still debate concerning the use of piperacillin-tazobactam to treat infections with ESBL-producing Enterobacteriaceae. There is some evidence that outcomes may be worse if the piperacillin-tazobactam MIC is 8 - 16 mg/l, with suggestions that if piperacillin-tazobactam is used to treat organisms with these MICs, prolonged infusions or more frequent dosing may be needed.[16]
The 87.4% susceptibility to cefoxitin suggests that plasmid-mediated AmpC enzymes are not particularly prevalent in this sample. The disparity in ciprofloxacin and levofloxacin resistance (21.7% difference) is intriguing and although we do not have genotypic data to support it, it may represent an increase in quinolone resistance mechanisms (e.g. plasmid-mediated mechanisms; enzymatic modification) other than the conventional target site mutations, where one would expect cross-resistance[171 as 61.5% ESBL of K. pneumoniae isolates were resistant to ciprofloxacin.
Among all aminoglycosides, amikacin exhibited the best activity, although this must be interpreted with caution as the stipulated breakpoints are high considering that this is a concentration-dependent agent. The MIC50 and MIC90 of 8 µg/ml and 16 µg/ml, respectively, indicate that for many isolates the probability of achieving adequate serum levels is quite low.
Overall, the MIC50 and MIC90 to most agents have not changed over the 3-year period, although the ability to detect smaller differences may have been limited by the methodological system employed for susceptibility testing. The one exception was colistin, for which a significant increase in MIC90 was noted in 2012 (p=0.001) (Table 4). This is alarming, as colistin represents the mainstay of the treatment of most of our extensively drug-resistant Gram-negative pathogens (including carbapenem-resistant K. pneumoniae)[18] Trends in susceptibility showed a significant decrease over the study period for most antimicrobial agents, with a few exceptions, e.g. piperacillin-tazobactam, levofloxacin and tigecycline.
A limitation of this study was that comprehensive patient demographic details were unavailable, and it was not possible to determine accurate trends in patient age, ward and gender. This information will be obtained in future surveys, allowing for more detailed analyses of antimicrobial resistance patterns, and how they change over time.
These surveillance results should be used together with hospital-specific data as tools in antimicrobial stewardship programmes and for the development of empiric therapy guidelines for healthcare-associated infections. Ongoing antimicrobial resistance surveillance, performed in a systematic and standardised manner, could be used as a tool to monitor the effectiveness of antimicrobial stewardship programmes that have been implemented in various centres across the country. Moreover, it provides a foundation for the systematic surveillance of important hospital pathogens that could be expanded and enhanced over time.
Conclusion
SA appears to have a relatively high percentage of ESBL-producing K. pneumoniae isolates in comparison with other geographical regions, which is of great concern, and a significant increase in ertapenem resistance over the surveillance period. This study presents the antibiotic resistance patterns of invasive K. pneumoniae isolates and gives an indication of the prevalence of resistance genes. Our findings provide important baseline data for further site-specific analysis of K. pneumoniae isolates, as well as a platform for enhanced surveillance of K. pneumoniae antimicrobial resistance in the country. Additionally, when analysed in conjunction with patient demographic and clinical details these data are important for the development of empiric therapy guidelines for management of sepsis in SA healthcare institutions.
Acknowledgements. This study was supported by GERMS-SA (the Group for Enteric, Respiratory and Meningeal disease Surveillance in South Africa). We thank Ms Serisha Naicker, Mr Elias Khomane, Mrs Zazi Molebatsi and Mrs Marshagne Smith for assistance with the laboratory work and Ms Penny Crowther for assistance with our database.
References
1. Ashley EA, Lubell Y, White NJ, et al. Antimicrobial susceptibility of bacterial isolates from community acquired infections in Sub- Saharan Africa and Asian low and middle income countries. Trop Med Int Health 2011; 16 (9):1167-1179. [http://dx.doi.org/10.1111/j.1365-3156.2011.02822.x] [ Links ]
2. Gutkind GO, Di Conza J, Power P, et al β-lactamase-mediated resistance: A biochemical, epidemiological and genetic overview. Curr Pharm Des 2013; 19(2):164-208. [http://dx.doi.org/10.2174/138161213804070320]
3. Brink A, Moolman J, da Silva MC, et al Antimicrobial susceptibility profile of selected bacteraemic pathogens from private institutions in South Africa. S Afr Med J 2007;97(4):273-279. [ Links ]
4. Brink AJ, Botha RF, Poswa X, et al Antimicrobial susceptibility of Gram-negative pathogens isolated from patients with complicated intra-abdominal infections in South African hospitals (SMART Study 2004-2009): Impact ofthe new carbapenem breakpoints. Surg Infect (Larchmt) 2012; 13(1):43-49. [http://dx.doi.org/10.1089/sur.2011.074] [ Links ]
5. Essack SY, Hall LM, Pillay DG, McFadyen ML, Livermore DM. Complexity and diversity of Klebsiella pneumoniae strains with extended-spectrum beta-lactamases isolated in 1994 and 1996 at a teaching hospital in Durban, South Africa. Antimicrob Agents Chemother 2001; 45(1):88-95. [http://dx.doi.org/10.1128/AAC.45.1.88-95.2001] [ Links ]
6. Paterson DL, Hujer KM, Hujer AM et al. Extended-spectrum beta-lactamases in Klebsiella pneumoniae bloodstream isolates from seven countries: Dominance and widespread prevalence of SHV- and CTX-M-type beta-lactamases. Antimicrob Agents Chemother 2003;47(11):3554-3560. [http://dx.doi.org/10.1128/AAC.47.11.3554-3560.2003] [ Links ]
7. Ehlers MM, Veldsman C, Makgotlho EP, et al. Detection of blaSHV, blaTEM and blaCTX-M antibiotic resistance genes in randomly selected bacterial pathogens from the Steve Biko Academic Hospital. FEMS Immunol Med Microbiol 2009; 56(3):191-196. [http://dx.doi.org/10.1111/j.1574-695X.2009.00564.x] [ Links ]
8. Performance Standards for Antimicrobial Susceptibility Testing: Twenty-Second Informational Supplement. Clinical and Laboratory Standards Institute. M100-S22 (Vol. 32, No. 1): 2012:45-61. [ Links ]
9. The European Committee on Antimicrobial Susceptibility Testing (EUCAST). Breakpoint tables for interpretation of MICs and zone diameters. http://www.eucast.org (accessed 3 July 2014). [ Links ]
10. Birkett CI, Ludlam HA, Woodford N, et al Real-time TaqMan PCR for rapid detection and typing of genes encoding CTX-M extended-spectrum beta-lactamases. J Med Microbiol 2007;56(1):52-55. [http://dx.doi.org/10.1099/jmm.0.46909-0] [ Links ]
11. Centers for Disease Control and Prevention. Multiplex real-time PCR detection of Klebsiella pneumoniae carbapenemase (KPC) and New Delhi metallo-p-lactamase (NDM-1) genes. 2012.http://www.cdc.gov/HAI/settings/lab/kpc-ndm1-lab-protocol.htm (accessed 3 July 2014). [ Links ]
12. Bamford C, Bonorchis K, Ryan A, et al. Antimicrobial susceptibility patterns of selected bacteraemic isolates from South African public sector hospitals, 2010. South African Journal of Infectious Diseases 2011;26(4):243-250. [ Links ]
13. Braykov NP, Eber MR, Klein EY, et al. Trends in resistance to carbapenems and third-generation cephalosporins among clinical isolates of Klebsiella pneumoniae in the United States, 1999-2010. Infect Control Hosp Epidemiol 2013;34(3):259-268. [http://dx.doi.org/10.1086/669523] [ Links ]
14. Mendes RE, Mendoza M, Banga Singh KK, et al. Regional resistance surveillance program results for twelve Asia-Pacific nations (2011). Antimicrob Agents Chemother 2013;57(11):5721-5726. [http://dx.doi.org/10.1128/AAC.01121-13] [ Links ]
15. Lowman W, Sriruttan C, Nana T, et al. NDM-1 has arrived: First report of a carbapenem resistance mechanism in South Africa. S Afr Med J 2011;101(12):873-875. [ Links ]
16. Nguyen HM, Shier KL, Graber CJ. Determining a clinical framework for use of cefepime and p-lactam/β- lactamase inhibitors in the treatment of infections caused by extended-spectrum-p-lactamase-producing Enterobacteriaceae. J Antimicrob Chemother 2014;69(4):871-880. [http://dx.doi.org/10.1093/jac/dkt450] [ Links ]
17. Robicsek A, Strahilevitz J, Jacoby GA, et al. Fluoroquinolone-modifying enzyme: A new adaptation of a common aminoglycoside acetyltransferase. Nat Med 2006; 12(1):83-88. [http://dx.doi.org/10.1038/nm1347] [ Links ]
18. Visser Kift E, Maartens G, Bamford C. Systematic review of the evidence for rational dosing of colistin. S Afr Med J 2014; 104(3):183-186. [http://dx.doi.org/10.7196/SAMJ.7011] [ Links ]
 Correspondence:
Correspondence:
O Perovic
olgap@nicd.ac.za
olga.perovic@nhls.ac.iza